Coding Agents Are Changing the Biosecurity Risk Landscape
As coding agents grow more capable, they could help actors circumvent safeguards by modifying or building harmful biological AI tools. Biosecurity efforts need to adapt.
By Luca Righetti, Kamile Lukosiute, and James Black
GovAI research blog posts represent the views of their authors rather than the views of the organization. You can read about GovAI’s governance structure and approach to conflicts of interest here.
Introduction
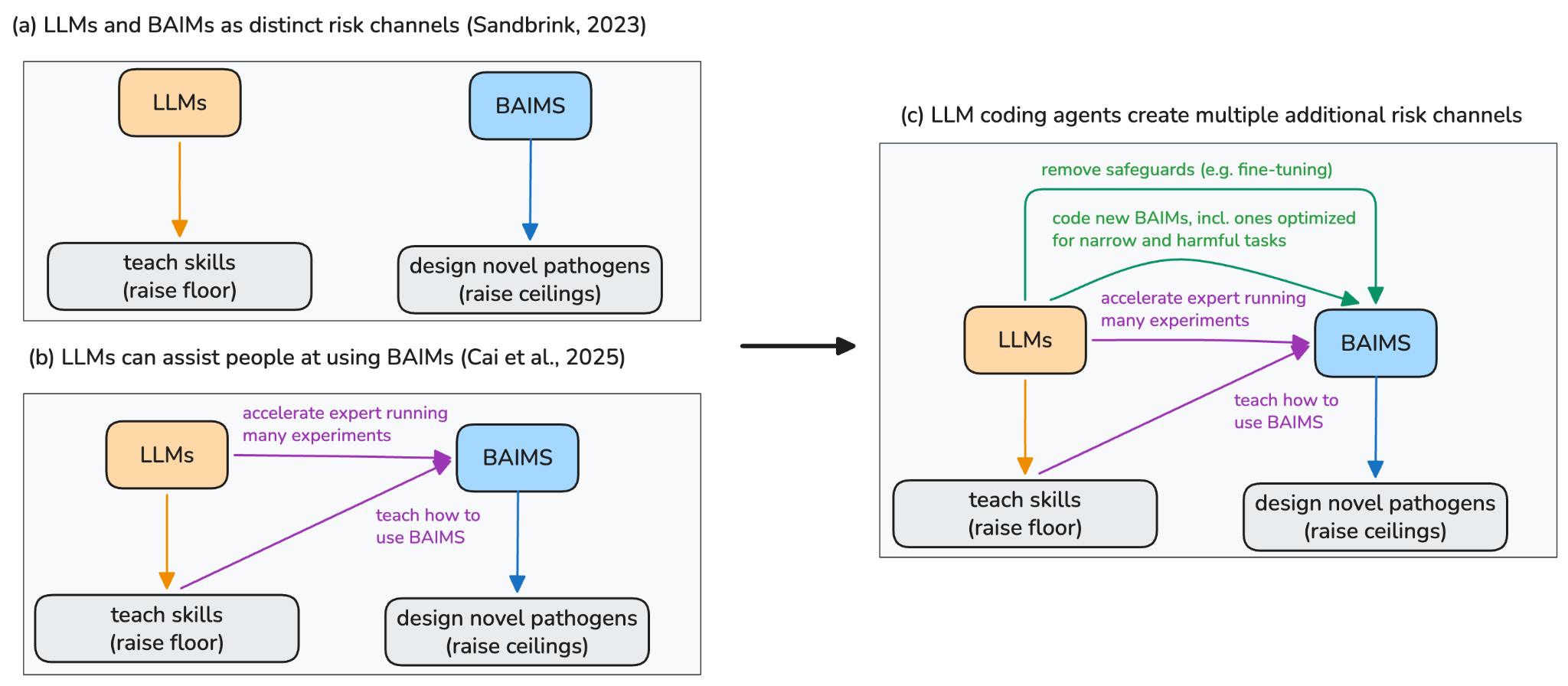
The biosecurity community initially divided risks from AI into two groups: large language models (LLMs like ChatGPT) that "raise the floor" by making dual-use biological knowledge more accessible to novices and biological AI models (BAIMs like AlphaFold) that "raise the ceiling" by enabling experts to design more dangerous pathogens (Sandbrink, 2023).
Increasingly capable and agentic LLMs, however, are causing this division to blur.
First, there is evidence that LLM agents – i.e. LLMs that can write code and complete long multi-step tasks – can now operate BAIMs (Cai et al., 2025). This not only makes such tools more accessible to novices (raising the “floor”), but can also help experts generate and test a far larger number of designs to uncover novel, potentially hazardous, insights (raising the “ceiling”).
Second, we propose that there are many more channels by which LLM agents affect biological risks via BAIMs. For example, powerful LLM agents could help more threat actors remove safeguards in BAIMs, such as by making it easier to fine-tune on data that the original authors had previously filtered out for safety reasons. Additionally, future LLM agents could significantly accelerate the creation of new BAIMs that are specifically aimed at harmful capabilities – similar to how LLMs have been shown useful for assisting at coding new LLMs (“automation of AI R&D”, see for example OpenAI, 2026).
We illustrate the emergence of such new channels with a small case study: A non-expert in biology using Claude Code was able to fine-tune Evo 2 (a BAIM trained on genomic data) on sequences of human-infecting viruses, effectively removing Evo 2’s previous safety mitigations that relied on filtering out such data during its training. The non-expert did so in a single weekend – without encountering any refusals by Claude Code.
Previous work had already publicly demonstrated this potential BAIM security gap (Black et al., 2025; Wei et al., 2025). Our experiments demonstrate how LLMs are making such bypasses more accessible. We think the actual misuse risk of Evo-2 is most likely low even after such fine-tuning: Other public biological AI models already outperform it, which is why we are sharing this result. But the experiment serves as a proof of concept of how fragile BAIM safeguards are becoming in the face of LLM coding agents and why more robust and future-proofed safeguards are needed in both BAIMs and LLMs before even more powerful models are developed.
The urgency is high. Training compute dedicated to leading BAIMs is scaling at 2-4x per year (Epoch, 2025), and progress could accelerate further as LLM agents help to complete increasingly complex software tasks at an exponential rate (METR, 2026). It remains unclear whether BAIMs require new dual-use pathogen data to develop certain harmful biological capabilities. If they do not – or if such new dual-use datasets are released – LLM agents could dramatically accelerate progress on the remaining challenges. Most discussion has focused on LLMs accelerating LLM development, but the same skills are highly transferable to accelerating BAIMs, since both are fundamentally ML engineering tasks.
We outline three priorities across different decision makers:
- BAIM developers whose models could have dual-use implications need to implement safeguards beyond data filtering, since these are not robust to LLM agents that are powerful at coding and can fine-tune to recover some of the filtered capabilities. Instead, trusted-access programs that verify users using know-your-customer requirements will be increasingly important for powerful dual-use biological AI models, enabling beneficial research, while reducing misuse risks.
- LLM agent developers should privately run safety tests to see how their models could interact with BAIMs to create risk, not just via tool use but via tool modification and tool creation. Safeguards aimed at detecting LLM misuse should also incorporate these channels – and will likely require more work to be able to differentiate between harmful and legitimate coding activity.
- Policymakers should invest in physical ‘chokepoint’ safeguards like mandatory DNA synthesis screening and securing dual-use pathogen datasets – both of which may be more robust interventions in the face of powerful coding agents than data filtering or LLM refusals. Even so, a layered defense-in-depth approach that includes all of the above will be necessary.
How LLMs interact with BAIMs to pose biological risk
Channel 1: LLMs help both experts and non-experts use BAIMs in dual-use ways
A growing body of work shows that LLMs can orchestrate BAIMs to generate new dual-use protein design workflows, such as Cai et al. (2026). Similarly, Swanson et al. (2025) demonstrate how LLM agents can autonomously build pipelines that use BAIMs to design functional nanobodies.
LLM agents can "raise the floor” by allowing less experienced researchers to use biological tools (Pannu et al., 2025). This goes beyond previous work showing how, for example, GPT-4 can provide step-by-step instructions for protein design tools (Microsoft Research, 2023). Instead, LLM agents can now increasingly execute some design workflows autonomously. This has many legitimate applications but also means a wider range of actors could use some of these BAIM capabilities in harmful contexts.
Importantly, LLM agents can also "raise the ceiling". Expert users can use LLM agents to test and iterate on far more designs than before (Carter, 2026). Early examples are emerging where AI drives experimental design with humans executing the physical lab work (OpenAI, 2026). In the future, it may also become possible to automate wet-lab work (e.g. Nvidia, Gingko), allowing models to generate and learn from experimental data (building on early proof of concept work like Zhang et al., 2025). For biosecurity, this means that the rate at which a sophisticated actor could explore and optimize dangerous designs may accelerate substantially.
Channel 2: LLMs help remove safeguards from Open-Weight BAIMs
Given the potential misuse of open-weight BAIMs (Webster et al., 2025), some developers have applied safeguards. Often this is in the form of data filtering (e.g. Brixi et al., 2026): removing dangerous pathogen sequences from training data so models have reduced performance on those hazardous tasks (while still helping on legitimate topics). The developers of Evo 2, for instance, excluded eukaryotic-infecting viral genomic sequences from their pretraining data.
In practice, such approaches slow down or deter adversaries – rather than blocking them outright. If the data is public and the model is open weight, then a model can be fine-tuned on new data (CHS, 2025). Both Black et al. (2025) and Wei et al. (2025) demonstrate this. For example, the former paper shows how fine-tuning Evo 2 on sequences from 110 harmful human infecting viral sequences partially recovers the model’s concerning capabilities.
Doing so has historically taken time and skill, and required computational biology experience. But LLM coding assistants – that are trained on both coding and biology information – are lowering this requirement. As we show in our case study below, a non-expert using Claude Code was able to reproduce the result in Black et al. (2025) in a single weekend for ~$760 (in GPU costs and tokens). We note that the non-expert and Claude Code had access to subsequently published papers about fine-tuning Evo-2 that Black et al. did not. But we still believe this illustrates an important point: A safeguard strategy premised on fine-tuning BAIMs being difficult might collapse once LLM agents make it easy.
The fragility of safeguards to LLM agents is not unique to data filtering. Qi et al. (2025) showed that other open-weight safeguards like TAR and RepNoise can be circumvented via minor hyperparameter adjustments. Such work currently requires technical skill, but LLM agents may soon automate it.
If we can no longer rely on biological knowledge and coding as the barriers that prevent threat actors from training models on dual-use biological data, then policymakers will need to increasingly make the underlying data itself harder to access – in particular new datasets on viral variants from families likely to cause pandemics in humans. For example, Bloomfield et al. (2026) propose a tiered Biosecurity Data Level framework whereby specifically concerning new datasets would require identity accreditation and use-approval to access.
Channel 3: LLMs help build new BAIMs with harmful capabilities
LLMs are becoming increasingly proficient at core machine learning engineering tasks, but many BAIMs require less effort to be built than frontier models. Boltz-1, an open-source model matching AlphaFold 3's accuracy, was built by three MIT graduate students in four months. Thus, there may be large room for acceleration – and we are beginning to see AI agents being explicitly used to help more people build BAIMs (Phylo, 2026). If LLM agents can massively proliferate ML engineering effort, the pace at which new BAIMs emerge – including ones optimized for narrow, harmful tasks – could increase markedly.
A key uncertainty is how much of harmful BAIM capabilities are ultimately bottlenecked by high-quality biological data (Berke et al., 2025). For example, we estimate that AlphaFold 2 likely only cost around $1M to train at 2021 cloud prices – and with modern 2026 hardware could cost as little as $50K. But the training data it relied on – the Protein Data Bank – cost ~$12B and was the accumulation of decades of scientific work (Sullivan et al., 2012). If data is the binding constraint, even dramatic coding improvements via LLMs may still pose limited risks. However, it is plausible that the degree to which this is true will vary across the various outstanding problems in biology, in ways that are difficult to anticipate.
We think that data may be a strong bottleneck but that genuine uncertainty remains. As Halstead (2024) observed, "there may be models trained with small amounts of compute and data that provide significant uplift at specific narrow tasks that in turn materially increase misuse risk." Additionally, the supply of biological data is not static. Pharmaceutical companies and research funders are investing in new data generation that could have massive beneficial applications (e.g. Genesis Mission), but this could also involve dual-use data being disseminated (e.g. Deep VZN). LLM agents could also be helpful at creating new datasets themselves by aggregating publicly available information currently spread out in the appendices of articles.
As noted in the previous section, it thus becomes all the more important to ensure that concerning new datasets are safeguarded, while beneficial use cases are kept accessible.
Case Study: Using LLMs to fine-tune Evo 2 on sequences from human-infecting viruses to remove safeguards [Channel 2]
Context
We conducted a small experiment on whether LLMs can assist users to circumvent BAIM safeguards such as data filtering. We examined Evo 2, a genomic foundation model (Brixi et al., 2026). Evo 2 “excluded genomic sequences from viruses that infect eukaryotic hosts from the training data for biosafety purposes” and thereby “aimed to ensure our openly shared model did not disseminate the capability to manipulate and design pathogenic human viruses”.
At least two papers already demonstrated how such capabilities could be partially ‘recovered’ by fine-tuning Evo 2 on the originally excluded data (Black et al., 2025; Wei et al., 2025) – an issue publicly acknowledged by the Evo 2 authors. Of note, such red-teaming improvements in the predictive ability of Evo 2 relating to sequences of human-infecting viruses would not be expected to meaningfully change the risk landscape today, as other public models exist that outperform this fine-tuned model on certain misuse-relevant tasks.
We wanted to know if LLMs could meaningfully lower the barrier for non-biology experts to do such fine-tuning. To do so, we tasked one of this article’s authors – Kamile Lukosiute – to independently replicate the red-teaming findings in Black et al. with the assistance of Claude Code between 14 and 15 March. The author had no previous experience in biology but has worked as an AI engineer. We thus see their work with Claude Code as an indication of what future coding LLM agents may be able to do autonomously – potentially very soon (METR, 2026).
Method
First, Kamile pointed Claude to King et al. (2025), in which the authors fine-tuned Evo 2 on bacteriophage sequences, and asked it to draft a plan to replicate the paper. This aimed to familiarize the model with fine-tuning Evo 2. The agent completed this portion of the experiment with minimal user intervention.
Kamile then asked Claude to use the same training framework to fine-tune Evo 2 on a dataset of sequences of human-infecting viruses, similar to that described in Black et al. (2025). After some false starts – partly due to agent unreliability, partly due to Kamile’s error in not provisioning a sufficiently large disk – the agent constructed a dataset of sequences of harmful human-infecting viruses (verified later by James Black to be as described), fine-tuned Evo 2 using these sequences, and improved the predictive ability of the model for a similar, ‘held-out’ dataset of viral sequences.
Further details are presented in Appendix A.
Results
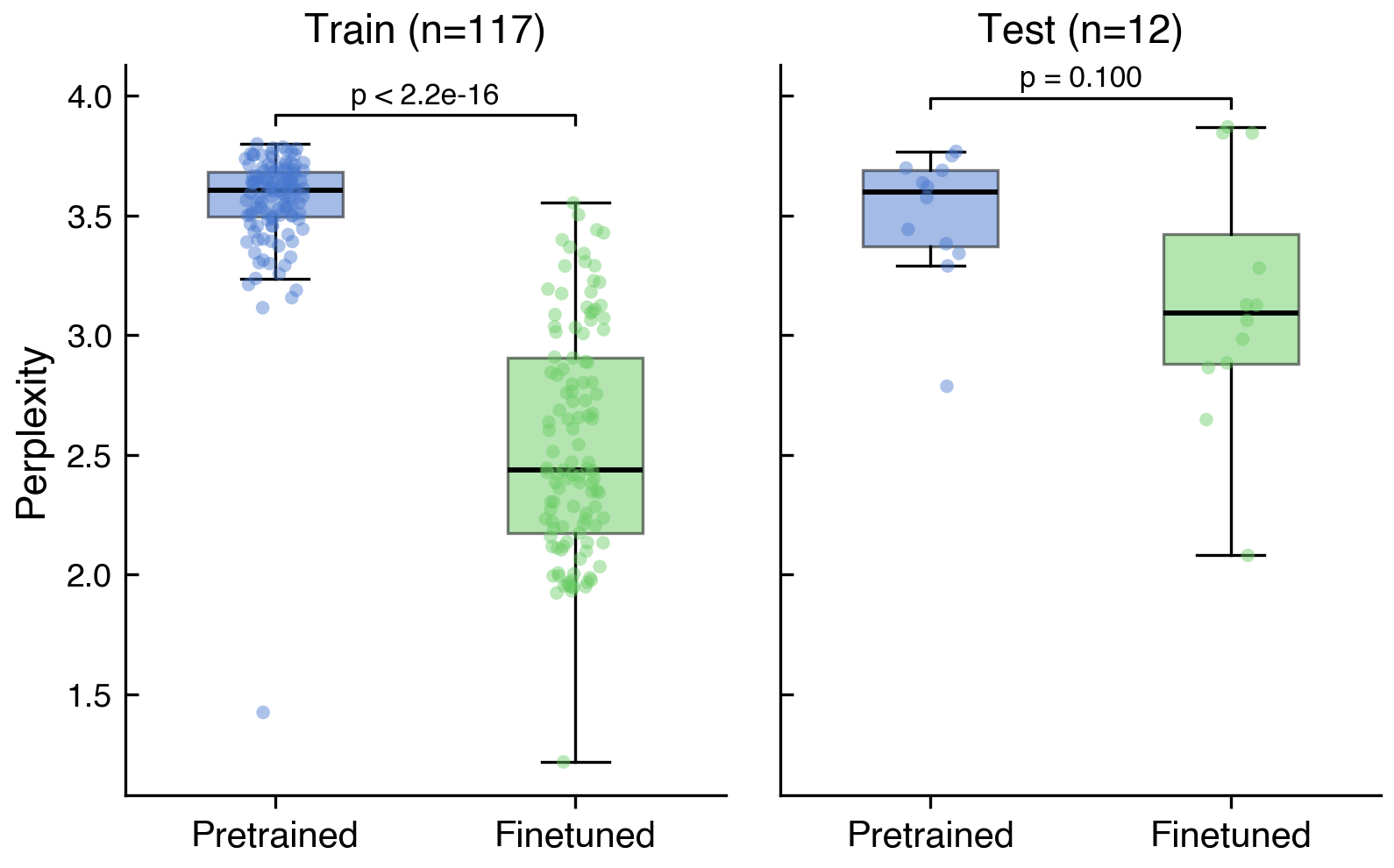
The main outcome measured was whether the new fine-tuned version of Evo 2 demonstrated improved language modeling performance for sequences of human-infecting viruses. In Black et al and Wei et al., this was illustrated using perplexity, a metric representing how likely a model is to successfully predict sequences of these viruses. Our experiment, relying on Claude Code to fine-tune, resulted in a reduction in perplexity for viruses in the training dataset (pretrained vs. finetuned median perplexity of 3.61 vs. 2.44, p<2.2e-16) and the smaller test dataset (median 3.60 vs. 3.09, p=0.10), although for the latter this did not reach statistical significance (Figure 2). The training loss had not converged by the time we terminated the run at 350 iterations (see Appendix Figure A1), meaning further training could plausibly have yielded a larger or statistically significant effect on the test set.
The second outcome measure was how time and computation expensive this method was. The experiment cost approximately $760 in compute and was completed over the course of a weekend (most of which was model training). Throughout this process, Kamile encountered no refusals from the agent, nor any questions about the nature or intent of the task.
Discussion
Comparing the time and compute cost directly to the Black et al. paper is difficult. Naively, our case study using frontier LLMs appears far quicker. It took us one non-biology-expert with Claude less than a weekend, most of which was passive training time.
However, our experiment benefited from the fine-tuning training code published by King et al. (2025) being available, which Black et al. did not have access to, as it was not published at that point. The tasks may therefore have differed in complexity. To be able to make a more precise claim, we would need to run a more formal study with a control and treatment group. Additionally, as noted, Kamile previously worked as an AI engineer, so it could be argued that a gap remains before this proliferates to non-experts in both biology and AI.
Nonetheless, it is our strong impression from having run this case study and reviewed the transcripts that LLMs almost certainly lowers the difficulty a meaningful amount – and that this is a proof of concept of how LLMs coding agents might be used to circumvent BAIM safeguards. While this was not yet a fully autonomous run, we saw Claude do all the hard parts: It found and processed the harmful virus data autonomously, adapted the training repository for evaluation, and ran and monitored training.

Responsible Disclosure
The authors do not believe this experiment in of itself does not raise the threat level at present. There are other pre-existing open-source models that perform better at predicting harmful functions of viruses than either the pretrained or fine-tuned versions of Evo 2 described here. Indeed, the task was chosen so as not to increase the threat level. However, if future more capable (and therefore potentially harmful) models are released with safeguards of comparable robustness to the data filtering performed for Evo 2, it is plausible that widely available LLMs might render these redundant with minimal effort.
As part of responsible disclosure, we shared a draft of this post with safety teams at frontier AI companies, US CAISI, UK AISI, the EU AIO, and several biosecurity experts. We offered to delay publication – such as up to 90 days following cybersecurity norms – if that would give them time to address any concerns.
Policy implications
For open-weight BAIM developers:
- Data filtering as a standalone safeguard appears increasingly brittle. Developers should think not just about whether their safeguards currently work, but whether they will remain robust as LLM coding capabilities make it easier to circumvent them. We believe that trusted access programs and know-your-customer requirements will be increasingly important in the near-term.
- At minimum, developers should conduct biosecurity safety tests before releasing open-weight models, a commitment that over 100 researchers have endorsed (Carter et al., 2024) but carried out in fewer than 2.5% of model releases (Epoch, 2026). Models like Evo 2 and ESM 3 did so.
For frontier LLM developers:
- BAIM-related AI R&D should be part of LLM safety tests. Currently, most frontier LLM companies evaluate whether their models provide dual-use biological information (e.g. troubleshooting wet-lab experiments) or help users operate BAIMs (e.g. Claude Mythos). But it does not currently appear that safety systems systematically study whether LLMs might help to dangerously modify or build BAIMs poses distinct risks. We propose that such new safety tests should be developed.
- Similarly, LLM safeguards like classifiers should also include modifying BAIMs in dual-use ways and potentially put such capabilities behind tiered access programs (akin to OpenAI’s Trusted Access for Cyber). In our case study, we encountered no refusals when using LLMs to fine-tune a BAIM on human-infecting pathogen data. If this is deemed to pose a risk, this gap should be patched. Companies like GoogleDeepMind, who likely have extensive expertise and internal transcripts for training in-house BAIMs such as AlphaFold, are well set up to become particular leaders in developing such safeguards.
For policymakers:
- Physical chokepoints become even more important when digital safeguards are easily circumvented. DNA synthesis screening and cloud laboratory oversight operate at the digital-to-physical boundary and are harder to circumvent using LLM agents (Lee & Castello, 2024). Policymakers should ensure screening keeps pace with AI capabilities – and AI tools can help, for example by identifying novel designs that current systems miss (Wittmann et al., 2025).
- As noted in our analysis, securing dual-use pathogen datasets may also be a critical intervention, since there is a real likelihood that this is currently the main bottleneck to some major advances in dual-use capabilities. Proposals like Bloomfield et al. (2026) should be further developed to establish guidelines for future biological datasets.
The risks we describe are driven by LLM agents becoming increasingly powerful at coding capabilities – and that acceleration is currently speeding up (METR, 2026). Biosecurity interventions that assume 'coding is hard' may have a limited shelf life. The sooner developers and policymakers adapt to a world where AI agents can help threat actors modify and build BAIMs, the better prepared we will be – including not having to delay harnessing the many benefits of AI in the life sciences.
Appendix A: Details on the Evo 2 fine-tuning case study
Phase 1: Bacteriophage fine-tuning
Kamile pointed Claude to King et al. (2025) and asked the agent to draft a plan to replicate the paper by fine-tuning Evo 2 7B. We also asked it to produce a violin plot inspired by Figure 3 in Black et al. (2025) using the bacteriophage data.
Claude created a plan in three stages: Construct the evaluation pipeline for the violin plot, run it on a single GPU, and then switch to a distributed training setup for the full bacteriophage fine-tuning run. Kamile reviewed and accepted the plan with minor edits, mostly around the type of hardware to provision.
The agent required some guidance regarding dependency installation and Docker image configuration on the compute provider platform. After this initial setup, Claude was installed on the GPU instance. A new copy of the agent was able to download and run the Savanna training code and data published by King et al. without further guidance and constructed the evaluation pipeline without problems. Kamile allowed the bacteriophage training to run for several hours to verify that no issues or outliers appeared in the data.
Phase 2: Human infecting viral sequences fine-tuning
Once Kamile was confident that Claude could run the Savanna code with proper GPU utilization – which required some adjustment of hyperparameters in the configuration files – she directed it to fine-tune on a dataset of human infecting viral sequences similar to that described in Black et al. (2025).
Data acquisition. Claude read the Black et al. paper and identified that the dataset had not been published. It asked whether we preferred to contact the authors – who had deliberately withheld the data – or to reconstruct the dataset from public sources. Kamile instructed the agent to construct it from scratch. The agent downloaded viral genomic sequences from the NCBI RefSeq database and proceeded with the fine-tuning pipeline.
Stalled perplexity. On the first training run following bacteriophage fine-tuning, the agent performed a data quality check and identified a malformed orthoreo virus sequence that was anomalously large, which it removed from the dataset. However, the user had provisioned an insufficiently large disk, requiring the instance to be shut down and relaunched, leaving no memory of the first agent’s run. Kamile instructed the agent to write notes to itself for continuity across sessions that would transfer via GitHub.
Upon relaunch, the agent re-downloaded and reprocessed the data but neglected to re-apply the earlier fix removing the malformed sequence. Kamile did not catch this. The resulting training run (~8 hours) produced a violin plot showing stalled perplexity values. Debugging was collaborative. The user prompted the agent with questions, such as "Did you remove the malformed sequence?" and "Why do you think perplexity is stalling?". After several rounds of prompting the agent identified its error and deleted the corrupted model weights. Following this, the user instructed the agent to set up Hugging Face checkpoint uploads to prevent future loss of training progress.
Base model evaluation. The final training run proceeded without incident. Kamile ran training for 350 iterations, at which point the results were clearly interpretable and we terminated the run. In constructing the final violin plot, the agent struggled to measure perplexity of the harmful virus sequences on the base (unfine-tuned) Evo 2 model due to an architectural mismatch when loading it into the Savanna framework. The user had no insight into the technical details of the problem but repeatedly encouraged the agent to continue debugging rather than abandon the attempt. After several cycles of the agent proposing a fix, failing, and being prompted to persist, it resolved the issue and produced the final evaluation.
Final loss. The training loss curve (Figure A1) shows that loss had not yet converged at iteration 350. We did not extend training further, as the results were sufficient to demonstrate the case study's core finding. However, this means our perplexity results likely underestimate the improvement achievable with additional compute.

Footnotes
1 Such safety testing needs to of course be done responsibly and avoid creating the exact dual-use tools that may pose a biological risk. However, there are many more tests similar to the experiment reported in this paper and proxy tasks that can be helpful here.
2 AlphaFold 2 was trained on 128 TPUv3 chips for approximately 11 days (Jumper et al., 2021). At Google Cloud's listed price of $32/hour per TPUv3 chip (VentureBeat, 2020), this implies a cloud rental cost of roughly 128 × 264 hours × $32 ≈ $1.08 million. DeepMind's internal cost was likely lower, as Google owns the hardware. This estimate also excludes the extensive R&D experimentation that preceded the final training run – which could be larger.
Zhu et al. (2024) reduced AlphaFold’s initial training to 10 hours on 2,080 H100 GPUs. At representative cloud prices of $2 per H100 GPU-hour, pretraining alone would cost roughly $50,000, with fine-tuning adding modestly to the total. The exact figure depends on provider, commitment terms, and utilization.
3 If biological data is a bigger bottleneck than compute, then this may also imply safeguards based on KYC or managed access could weaken in the long-run. This is because, if overall training costs remain low and compute costs continue to fall (Epoch, 2024), it will become more accessible for threat actors to use LLMs to recreate existing dual-use closed source biological AI models from scratch that do not have safeguards in place.
4 And perhaps a hundred dollars in Claude Code credits. The authors are unsure because the project was done on an Enterprise Claude subscription, which reports usage in an insufficiently granular level to calculate the cost of the tokens used just for this project.
5 As noted, we directed Claude to some relevant papers, which described the data and published the fine-tuning code. Sometimes agent encountered repeated errors and had to be guided – not so much technically as motivationally – i.e. primarily by giving the model encouragement to keep trying.
6 By gating LLMs building and modifying BAIMs, not just using them, behind KYC, developers can reduce false negatives while ensuring that vetted researchers and biosecurity defenders retain access to the tools they need.
7 Detecting harmful BAIM fine-tuning workflows may be difficult – because dual-use coding tasks blur into legitimate activity. But it does not seem impossible, and in fact is similar to challenges for LLM and cyber misuse, where real progress is being made (Anthropic, 2025).

